Dear all,
I would like to offer up an interesting article for discussion. Titled “Evolution: like any other science it is predictable”, it is written by Simon Conway Morris, professor of evolutionary palaeobiology at Cambridge. I’ve included the citation at the end of this post, and a free pdf version of the article is available here.
The whole article is worth a read, but I would like to focus my attention on two quotes in particular:
The case of the animals, in which we may take at least a parochial interest, is particularly problematic at present. Thus, so far as the fossil record is concerned, the Ediacaran assemblages appear to offer some tantalizing insights into the diversity of early metazoans (and quite possibly other distantly related macroscopic groups), but to date they give no clues as to the transition from more primitive forms. Molecular phylogenies are scarcely more helpful because, while there is strong evidence for the protistan choanoflagellates being the sister group of animals (e.g. Carr et al. 2008), neither they nor related groups that include corallochyteans, ichthyosporeans and ministeriids (e.g. Steenkamp et al. 2006) give many clues as to how the transformation to animals might have been achieved. Certainly, the prior existence of genes linked to cell adhesion and signalling in choanoflagellates (King et al. 2008; Ruiz-Trillo et al. 2008) points to the pre-adaptations for multicellularity. Note, however, that in at least one choanoflagellate, the unicellular(!) Monosiga, the tyrosine kinase signalling apparatus is not only far more diverse than any metazoan (Manning et al. 2008, p. 9678), but as the investigators note this network reveals ‘several common themes that suggest convergent evolution and a limited set of recurring molecular themes favoured by signalling pathways’. (My emphasis)
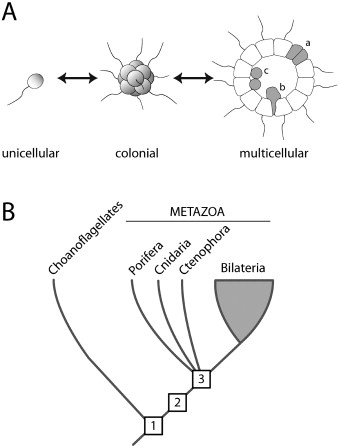
Choanoflagellates are small, single-celled eukaryotes that use their whip-like flagella to move through the water (these flagella are constructed in a way entirely different from the bacterial flagellum, and the two structures are not believed to be related).
As choanoflagellates are single-celled, they were not expected to contain genes for cell adhesion and signalling molecules. As Nicole King, the biologist who made the discovery, said in an interview: “I was surprised to learn that so much of animal biology was in place before the origin of animals.”
The fact that genes associated with multicellularity should predate multicellular organisms is consistent with, but not expected from conventional evolutionary biology. It is, however, just what we should expect if the first life on earth was engineered with its future evolution in mind.
Okay, but maybe that finding was just a fluke, you think. Not so. As Conway Morris writes:
Perhaps, one day, the entire molecular and morphological transition from protist to animal will be available, but if we enquire what fundamentally is required to make an animal among the most important presumably are: homeotic genes, structural molecules such as collagen, muscles for movement and nerves for the rapid propagation of information. Once again, it is difficult to see what might have prevented them from evolving. Thus, the homeodomain (HD) proteins go deep into eukaryotic history, and their presence in metazoans, fungi, Dictyostelium and plants points to a role in the evolution of multicellularity. Despite this striking association, Derelle et al. (2007) also argued that the last common ancestor of all eukaryotes possessed at least two types (TALE, non-TALE) of HD protein and, echoing the story of the SNARE genes, suggest that the rounds of duplication occurred independently. This leads them to ‘suggest that the eukaryotes as a whole are pre-adapted for multicellularity’ (Derelle et al. 2007, p. 217). In the context of evolutionary likelihoods, if not inevitabilities, it is also important to note that striking structural analogues to the HD proteins occur in the prokaryotes (Treisman et al. 1992; Kant et al. 2002). The independent emergence of more complex homeotic systems does not seem that improbable. (My emphasis)
Homeodomain proteins play a vital role in the development of multicellular organisms, where they are considered part of the development toolkit. The researchers found that homeodomain proteins “are present in all eukaryotic lineages containing multicellular organisms, and absent in exclusively unicellular lineages.” In other words, homeodomain proteins does not appear to play an important role in unicellular organisms.
But the researchers carried out a phylogentic analysis and concluded that the multicellular organisms with homeodomain proteins (animals, plants, algae, and fungi) had all inherited them from a single-celled ancestor, the “Ur-eukaryota”, while those lineages who remained sigle-celled lost their copies through reductive evolution:
As a corollary of ancestral molecular complexity, Ur-eukaryota probably possessed many of the good building blocks, which were subsequently recruited, by convergence in several lineages, to perform the functions required for development of multicellular organisms. In other terms, we suggest that the eukaryotes as a whole are preadapted for multicellularity, which only means that the ancestral complexity of the eukaryote genome and cell biology facilitated multiple acquisitions of multicellularity. (My emphasis)
“Preadapted for multicellularity”. Natural selection is the Blind Watchmaker, with no thought for anything that doesn’t increase the fitness of organisms right now. But thinking about the future, that smacks of teleology.
Thoughts?
References
Conway Morris S., 2010, “Evolution: like any other science it is is predictable”, Philosophical Transactions of the Royal Society B 365 (1537):133-145
Derelle R., et al., 2007, “Homeodomain proteins belong to the ancestral molecular toolkit of Eukaryotes”, Evolution & Development, 9(3):212-219
King N., 2004, “The Unicellular Ancestry of Animal Development”, Developmental Cell 7(3):313-325