Continuing the discussion from Responding to pastor's hesitations:
Before getting into the nitty gritty of endogenous retroviruses (ERVs), let’s start with a simple analogy to explain how the evidence works.
Let’s say you have two volunteers and two unabridged Oxford English Dictionaries. You give one dictionary to each person and tell them to go into separate rooms where they can’t see what the other is doing. You then ask them to randomly pick 100 words out of the dictionary and write them down on a sheet of paper. You bring them back into the same room and compare their list of words.
What would you expect? You would expect that few, if any, of the words should match. There are thousands and thousands of words in the OED and the chances of randomly matching nearly all 100 words from two random searches is highly improbable. As you would expect, none of the words match up. However, if you asked them to randomly find 1,000 words they may match a handful . . . maybe.
This is analogous to the way in which the ERV evidence works. In order for the retrovirus to replicate it has to insert its genome into the genome of the cell it has infected. Once in the host genome, the strong promoters in the viral genome cause the cell’s transcription and translation systems to make new viral particles from the genes in the viral genome. The process of inserting the viral genome into the host genome is quite random. If this happens in a gamete (i.e. egg or sperm), then it is possible for the viral insertion to be passed on to offspring. This is called an endogenous retrovirus, or ERV.
Due to the random nature of viral integration into the host genome we wouldn’t expect two independent insertion events to occur at the same position in two cells. The only reason we would expect to see large numbers of ERVs shared at the same position in two different genomes is if that insertion happened once in a common ancestor. The fact that you have about 200,000 ERVs in your genome that is shared by your siblings, extended family, and all other humans is testament to this process. Its not as if you and your cousin suffered 200,000 retroviral infections that just happened to produce the very same random insertions at the same spot in both of your genomes. The same logic applies to different species.
So how many ERVs do we share with other species? A lot of them. The human genome project found just over 200,000 ERVs in the human genome. When they sequenced the chimp genome they found that 82 of those human ERVs were not found at the same spot in the chimp genome. More than 99.9% of human ERVs were found at the same spot in the chimp genome. nearly all 200,000 of them. This couldn’t happen by random independent events. This can only happen by common ancestry.
References.
Human genome paper, table 11:
http://www.nature.com/nature/journal/v409/n6822/fig_tab/409860a0_T11.html
Chimp genome paper, table 2:
http://www.nature.com/nature/journal/v437/n7055/fig_tab/nature04072_T2.html
There is also evidence in the ERV sequences themselves that provide more independent lines of evidence for common ancestry, such as LTR divergence. These are bit more complicated and probably not conducive to the type of conversation you are having with your pastor. There are also many common creationist arguments which I could cover. Such as:
How do you know that ERVs are from retroviruses?
Is retroviral insertion random?
How do you know that God didn’t create these genomes with the ERVs already in them (i.e. the Omphalos argument)?
If any of these interest you I would be happy to discuss them, or answer any other questions that you may have.
Is there a typo here? If not, I have lost you! We are saying that they are incredibly similar aren’t we?
I just edited that part. Just to clarify, nearly all 200,000 human ERVs are found at the same position in the chimp genome. There are just 82 human ERVs not found at the same position in the chimp genome which are considered to be insertions that have occurred in the human lineage (i.e. species specific) since the chimp and human lineage split.
I think he is likely to believe me rather than asking these questions, but if there are simple answers, that would be very helpful!
Here is another resource with a lot of q/a regarding ERVs: Veritas: Endogenous RetroViruses - Frequently Asked Questions
Some of those are beyond my ability to understand! But I am glad those questions have been asked, and that there are well presented answers which appear to make sense!
Also, the fact that this is Veritas means it isn’t just an in crowd at BioLogos who are saying these things! If OCCA (Oxford Apologists) were to say anything about this, it would be even more convincing for my pastor, but I don’t think they do.
They answers aren’t necessarily simple, but they are interesting if you like these sort of things.
If we go back to the dictionary analogy, we can talk a bit about insertional hotspots. These are regions of the genome that retroviruses have a tendency for inserting into.
Let’s say we took those two lists of words that our two volunteers picked. We look up those words in the dictionary and plot the page number and their position on the page. What we will probably find is that our volunteers tended to pick words from pages towards the center of the dictionary. They also tended to pick words from the page on their dominant side, and words that were towards the center of the page. However, there are still thousands and thousands of words to randomly choose from in the middle section of the book, the middle sections of each page, and on the left or right page. Even then, these are only tendencies and our volunteers did still pick words that were outside of these “hotspots”.
This is somewhat analogous to insertional hotspots. Some retroviruses like to insert into areas that are being actively transcribed. Other retroviruses prefer to insert into regions that may have higher AT content or GC content. In the end, we are still talking about hundreds of millions to billions of bases that are found in the favorable regions, and these retroviruses still insert outside of these favorable regions.
Creationists will also argue that it makes sense that the same retrovirus will insert into the same position in genomes that are similar. This makes no sense once you actually look at the science. One such study worth looking at deals with three viruses: MLV, HIV. and ASLV. They allowed these viruses to infect cultured human cells that were genetically identical. What did they see? They saw that these viruses inserted all over the place. You can see it for yourself in Figure 1 in this paper:
In the results they state:
"For HIV the frequency of integration in transcription units ranged from 75% to 80%, while the frequency for MLV was 61% and for ASLV was 57%. For comparison, about 45% of the human genome is composed of transcription units (using the Acembly gene definition). "
So HIV tended to insert into transcription units (i.e. regions where there are more genes) to the tune of 75 to 80%. However, transcription units make up nearly 50% of the 3 billion base haploid genome. That’s 1.5 billion possible places for the virus to insert. The other 20% of the time it inserts into the other 1.5 billion bases.
Another favorite is this quote:
“There were identified `hot spots’ containing integration sites used up to 280 times more frequently than predicted mathematically.”
http://www.sciencedirect.com/science/article/pii/S0014579398004785#BIB41
What they don’t tell you is that the base probability was around 1 in 10 million, if memory serves. Increasing your odds from 1 in 10 million to 280 in 10 million can not explain why 99.9% of human and chimp ERVs are found at the same spot in each genome.
This got a bit long, but I hope that helps. If nothing else, it can give you some confidence that you are passing on solid information to your pastor.
Thank you! I do have that confidence and my pastor trusts my judgement! So I think this will be a very good example for him.
Or may I suggest the more simple and false, “Both sides are interpreting the same evidence.”
I am making notes of things to pass on to my pastor, and I have included a lot of the input above. I came up with my own analogy, and I thought I would share it in case it is helpful for anyone else. Some creationists have argued that because BioLogos accepts the idea of “random” in evolution, that they believe creation was lacking in purpose!
Here is my analogy:
Evolution includes certain random processes in a mathematical sense, but scientists are not assuming this means a lack of purpose. If you tip a bucket of water over my head, the particular drops of water fall in random places, but the end result is that I get wet! It would be crazy for me to argue that you don’t exist because you didn’t place every drop where it landed! My energy would be better spent crying out “What did you do that for?!”
Here is another analogy geared towards christians for you to ponder:
When scientists say that a process is random what they are saying is the observable results are indistinguishable from a random process. Science doesn’t test metaphysical claims, but rather tests hypotheses. If there is purpose that appears to be random then science wouldn’t be able to detect it.
As an analogy, we could look to Jonah:
Jonah 1:7 Each man said to his mate, “Come, let us cast lots so we may learn on whose account this calamity has struck us.” So they cast lots and the lot fell on Jonah.
Most of the sailors on that ship would have seen what appeared to be a random occurrence, but was there purpose behind it?
Your discussion of “random processes” will always be vulnerable to misinterpretation if you don’t include the usual disclaimer that God knows the difference between the “random appearance of some orderly things” and things that may satisfy the mathematic definition of randomness, but which are truly guided by his Will and Purpose.
And Proverbs 16:33 says “The lot is cast into the lap, But its every decision is from the LORD.” So yes there is purpose behind every “random” event.
That is even better than the verse I used. There appears to already be a tradition within Judeo-Christian theology that there is a deeper mystery underneath random events. Therefore, things like random mutations or random retroviral insertions shouldn’t be a problem for the theological position where God works through natural processes, at least from what I can see.
Thanks. That is true, but I doubt that my chat with my pastor will go to that level. He isn’t really into maths. The reaction to “random” if he has one is probably more of an emotional one brought on by ID people discussing it as if it means “aimless”. My tipping water over my head example is a bit of fun, but one that he will probably appreciate! Just so long as he doesn’t feel the need to have it as a sermon illustration with me as the victim!
In previous posts I discussed the position of ERVs in genomes, and how they evidence common ancestry between humans and other apes. Here is an excerpt from a 1999 peer reviewed paper by Johnson and Coffin:
“Given the size of vertebrate genomes (>1 × 109 bp) and the random nature of retroviral integration (22, 23), multiple integrations (and subsequent fixation) of ERV loci at precisely the same location are highly unlikely (24). Therefore, an ERV locus shared by two or more species is descended from a single integration event and is proof that the species share a common ancestor into whose germ line the original integration took place (14).”
http://www.pnas.org/content/96/18/10254.full
As with all good science, it is always a good idea to test a theory from multiple independent lines of evidence. ERVs actually supply 3 different lines of independent evidence for common ancestry, one of which I discussed in previous posts. The next line of independent evidence is LTR divergence. From Johnson and Coffin (1999):
“Third, sequence divergence between the LTRs at the ends of a given provirus provides an important and unique source of phylogenetic information. The LTRs are created during reverse transcription to regenerate cis-acting elements required for integration and transcription. Because of the mechanism of reverse transcription, the two LTRs must be identical at the time of integration, even if they differed in the precursor provirus (Fig. 1A). Over time, they will diverge in sequence because of substitutions, insertions, and deletions acquired during cellular DNA replication.”
Here is a diagram of a retroviral genome:
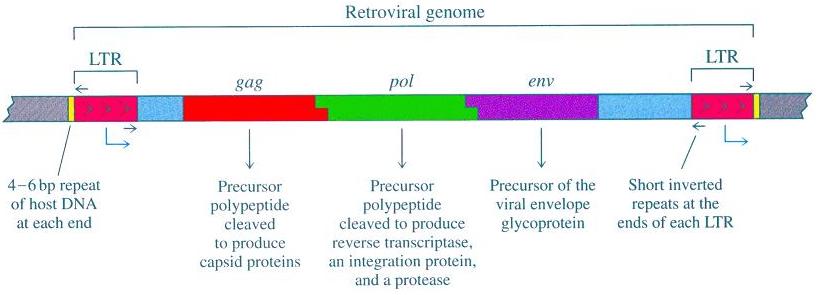
As you can see, the LTRs, or long terminal repeats, are like the bookends of the retroviral genome, called the 5’ and 3’ LTRs. As part of the retroviral life cycle, one LTR on one side is copied to produce the other LTR on the other side. Therefore, when the retrovirus inserts into the host genome the two bookends (the LTRs) have identical sequence. Once in the host genome, the slow accumulation of different mutations in each LTR will cause the LTRs to become less similar over time.
So how does this phenomenon allow us to test common ancestry? Evolution predicts that LTR sequences should reflect evolutionary distance, and that they should diverge from each other. That is, there should be more differences between the orangutan and human LTRs than between the chimp and human LTRS. If we use LTR sequences as inputs into phylogenetic algorithms then the phylogenies produced by these algorithms should match the phylogenies constructed by morphology (i.e. what the species look like). This is the standard phylogeny for apes:

From the Johnson and Coffin (1999) paper, this is the phylogeny of shared LTRs among different primate species:

(click on image for larger version)
As you can see, the standard phylogeny and the LTR phylogenies match for almost all nodes. On top of that, the 5’ and 3’ LTRs are on different branches of the phylogeny, consistent with the expected divergence. This is yet another independent line of evidence that screams common descent. Only evolution predicts a phylogeny of LTR sequences. ID/creationism does not.
Nice! I don’t understand every word, but I understand enough to see how good it is!
I’ve had two chats with my pastor now, and he has certainly taken some of the evidence on board. He also told someone else to look up Biologos!
He has asked me to prepare 6-10 weeks’ worth of material for a small group looking at resolving faith and science issues. He said he might come! That’ll be next year, so I might start a post then to see what ideas people have for what should go in the sessions. But I already have some good stuff now, so thanks everyone!
I always love to answer science related questions, so don’t be shy. I am always trying to improve how I present this info to the general public so your input would be very helpful. When you know the science in and out it can be difficult to judge how well you are getting across to people who don’t understand the science.